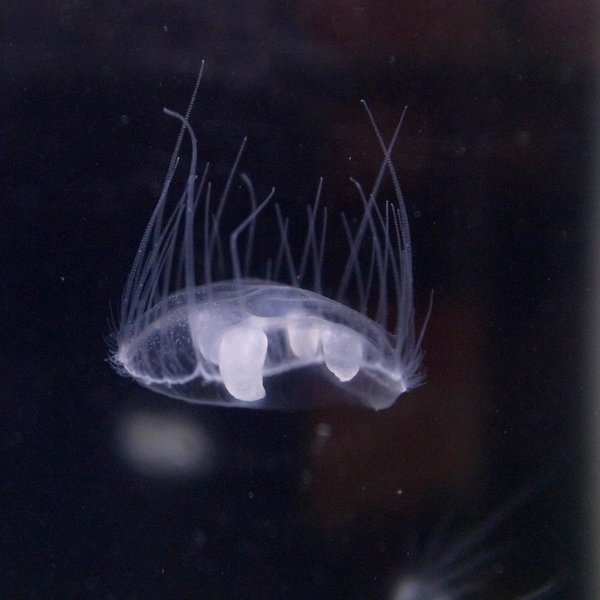
Craspedacusta sowerbii
A cryptic invasive spreading silently through Michigan's inland lakes. Population genomics could reveal how it colonizes new waterways and inform early detection strategies.
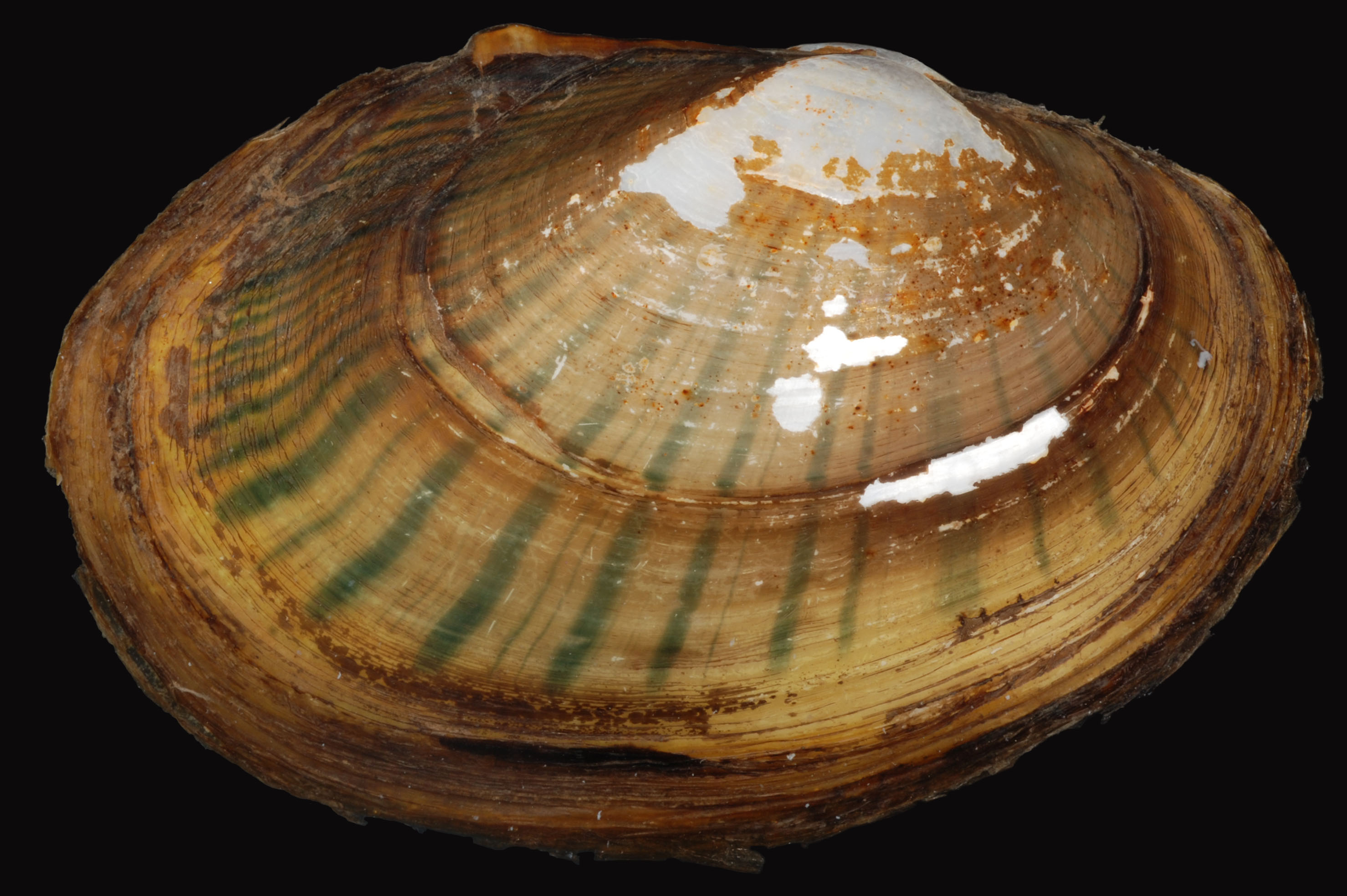

Lampsilis & Epioblasma
Michigan hosts 45 species of freshwater mussels — among the most endangered animals in North America. Most lack reference genomes critical for conservation management.
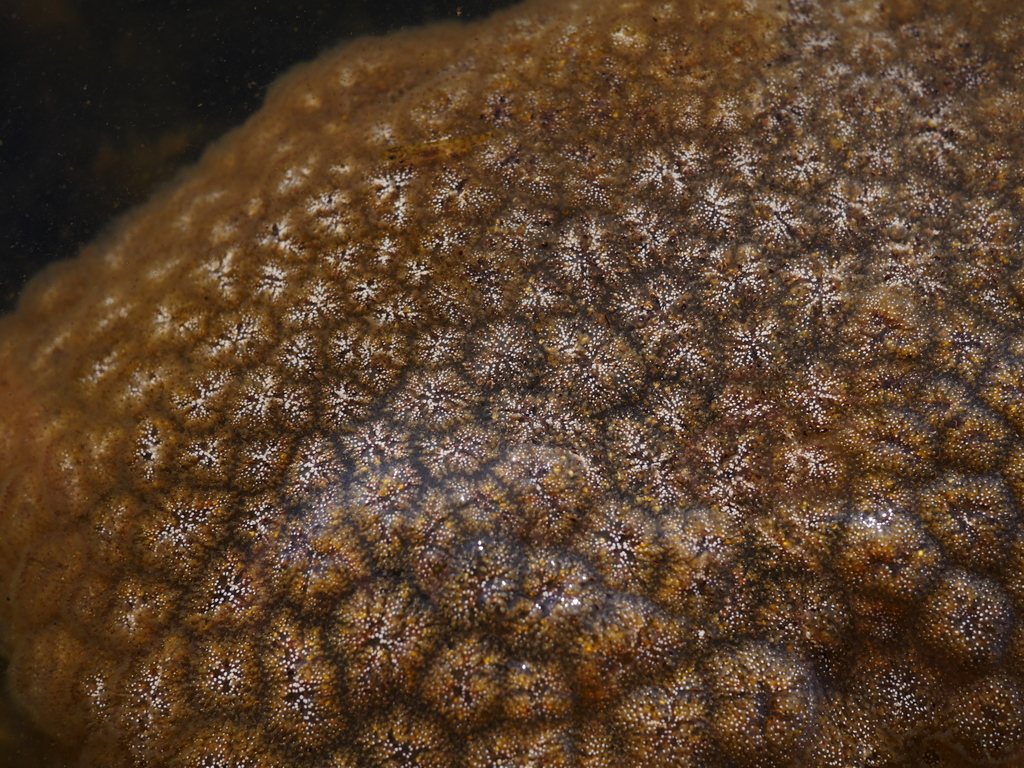

Pectinatella magnifica
Gelatinous, brain-like colonies that grow on submerged branches. People find them and have no idea what they're looking at. No nuclear genome exists — not even from its native range in North America.
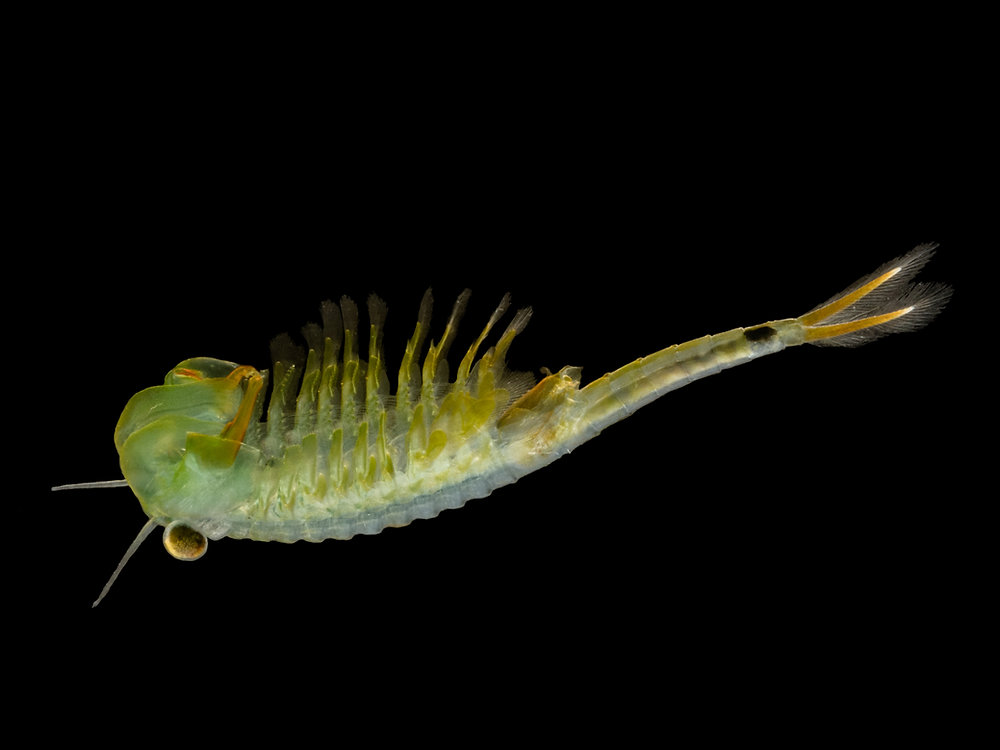

Eubranchipus bundyi
Ancient crustaceans that hatch in Michigan's vernal pools each spring and vanish weeks later. No genome exists for any of the 21 species in this genus.
More organisms coming — nominate a species